Hi, I'm Wesley DeMontigny. I build probabilistic models and scientific software.
Completed Work
Publication/Software
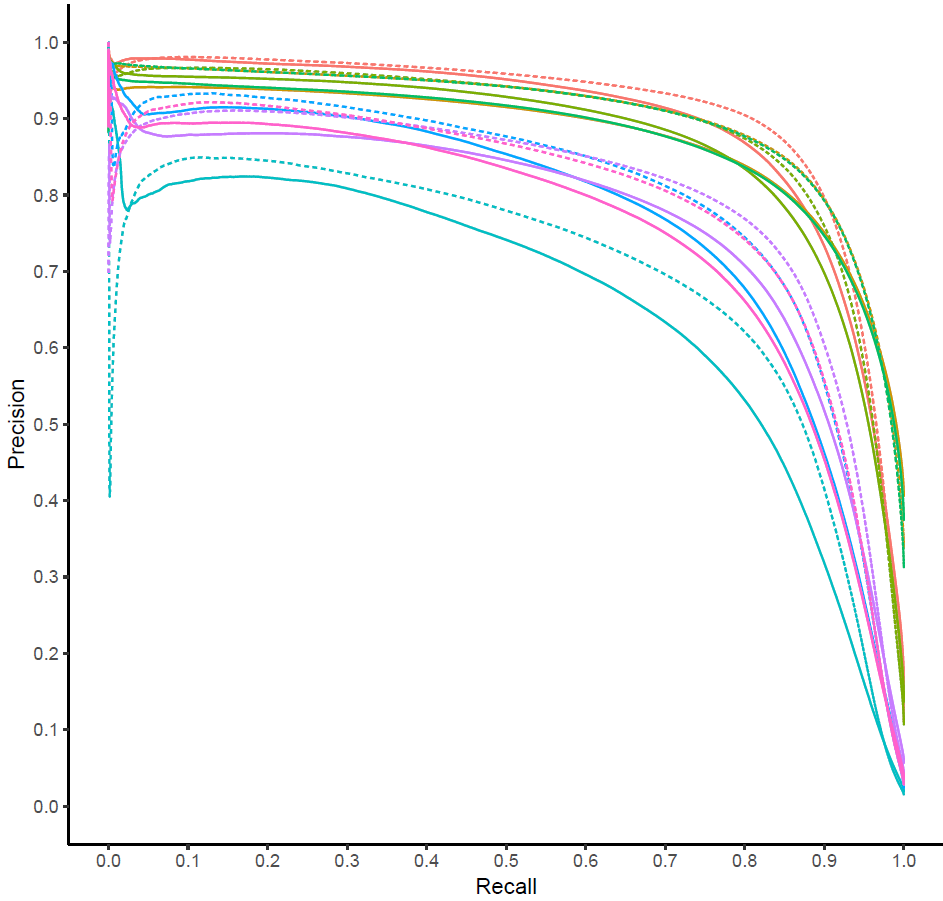
Inferring Gene Presence in Incomplete Data via Phylogenetic Occupancy Modeling
A probabilistic graphical model for imputing missing genes in large, highly-incomplete, genomic datasets.
Read the preprintOpen repository
Publication
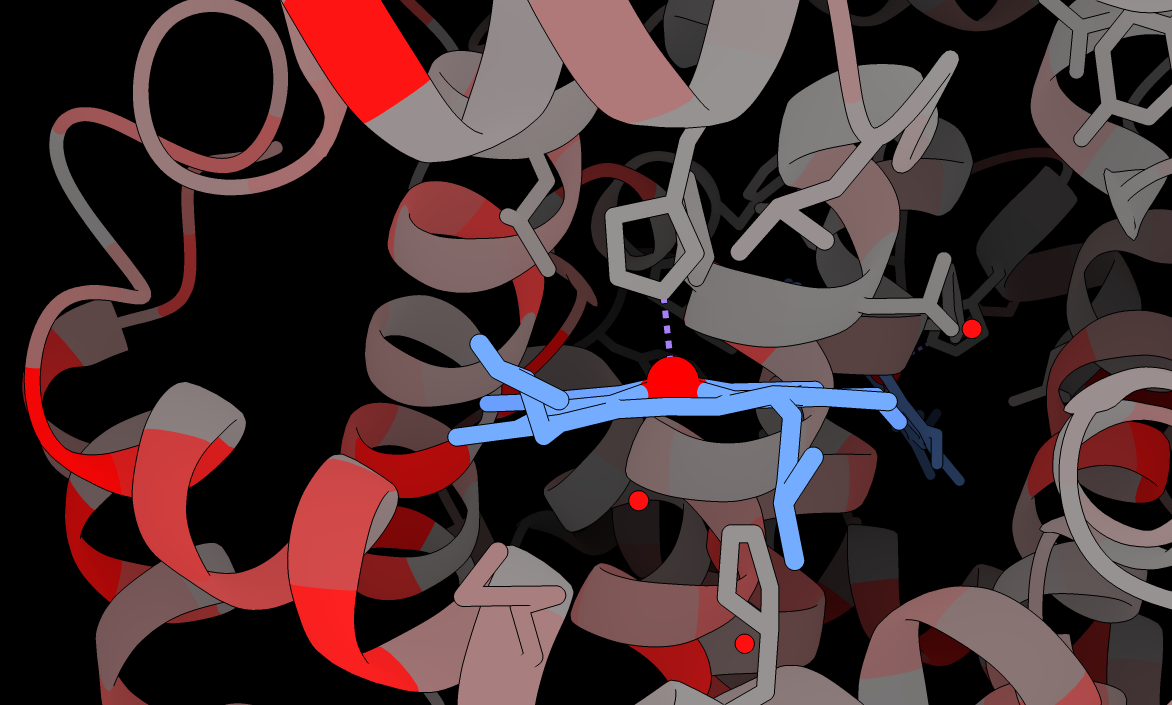
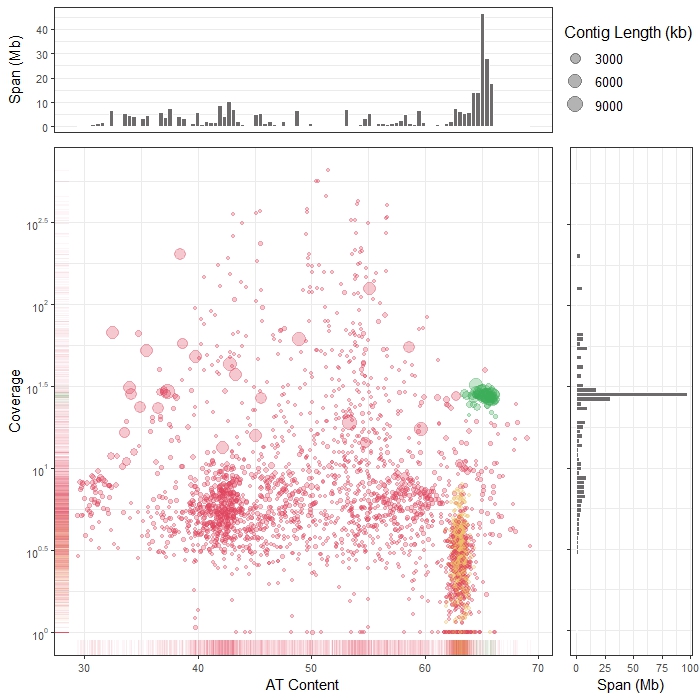
The nuclear and mitochondrial genomes of Amoebophrya
Bioinformatic analysis of a novel marine parasite genome, including alternative genetic code usage and heterogeneity of mitochondrial genome.
View publicationSoftware
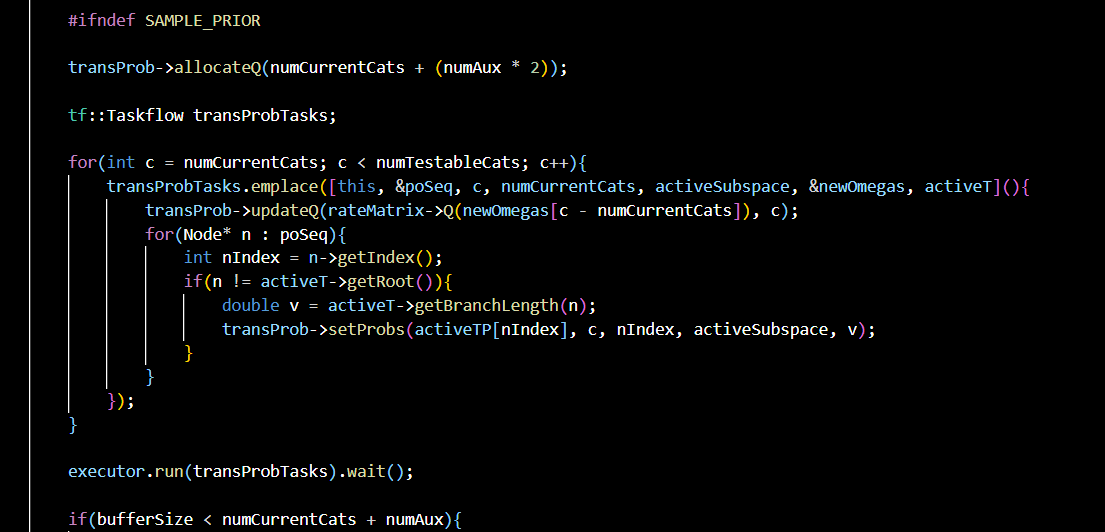
Smc-CAT
A C++ sequential Monte Carlo sampler for the CAT model, an important nonparametric Bayesian phylogenetic model.
Open repositoryMy Current Work
Machine learning
Graph neural networks for gene presence prediction
Graph neural networks for link prediction between genes and genomes, drawing inspiration from social recommendation systems.
Statistical modeling
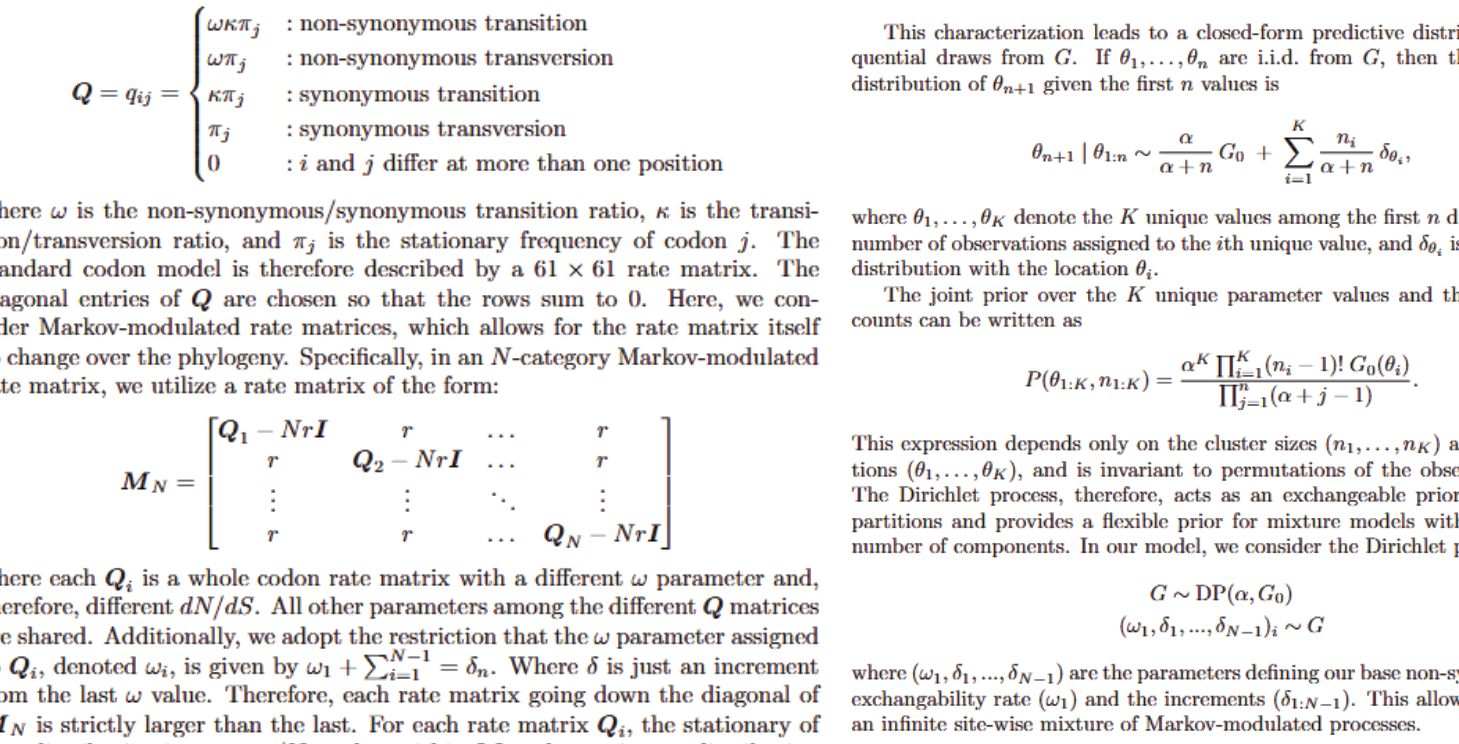
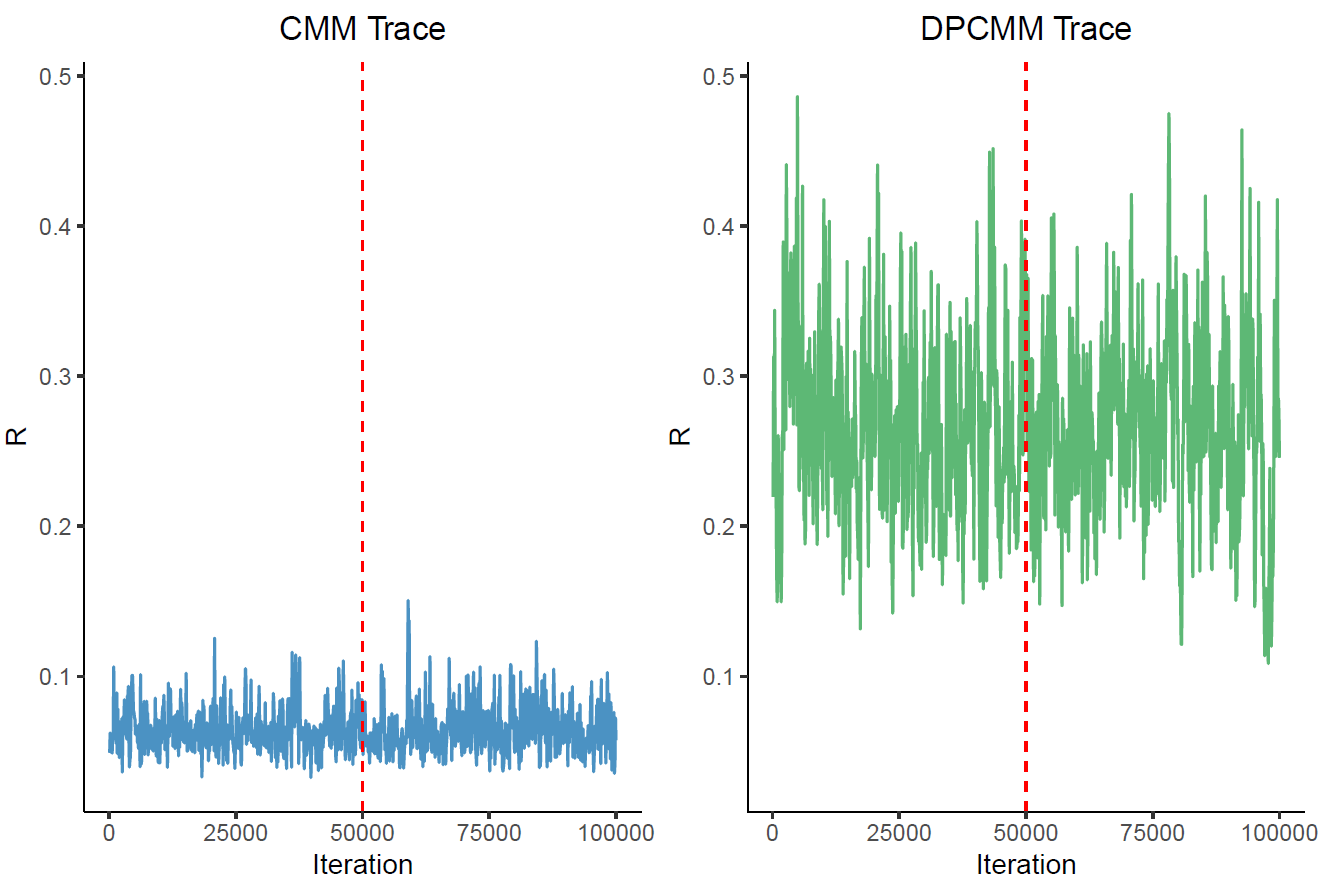
Reversible-jump MCMC for hidden evolutionary regimes
A C++ inference engine for infinite-mixture models that detects hidden regime changes in stochastic evolutionary processes.
Statistical modeling
Automatic partitioned phylogenetic analyses
A nonparametric Bayesian model that automatically partitions genomic sequences by shared evolutionary parameters.
Statistical modeling
Gene presence imputation with shared latent histories
A probabilistic clustering model that jointly infers shared gene histories to impute missing genes in partially observed genomes.